

On November 17, 2022, Professor Wang Xinghuan's team at Zhongnan Hospital of Wuhan University published the latest research results in Frontiers in Genetics, an important journal in the field of genetics, named "Genetically Supported Causality Between Benign Prostate Hyperplasia and Urinary Bladder Neoplasms: A Mendelian Randomization Study"[1].

This study used the two-sample Mendelian Randomization (MR) method to verify Benign prostate hyperplasia (BPH) and Bladder cancer (BLCA) at the gene level. There is a causal relationship, that is, BPH can increase the risk of BLCA, while BLCA cannot increase the risk of BPH.
The basic issues in traditional epidemiological research are "correlation" and "causal relationship". Clarifying the causal relationship between diseases is extremely important for guiding clinical work. At present, the "gold standard" for empirical testing of scientific hypotheses is conducting randomized controlled trials (RCT), but RCT has inherent flaws. For example, the results of observational studies are easily disrupted by confounding factors and it is difficult to obtain targeted interventions that only affect the target risk factors, resulting in causal relationships that are not actually established. For example, some articles reported that people who often "drink red wine" have a low incidence rate of coronary heart disease, but people who often consume red wine often live in regions with high "socio-economic level", so "socio-economic level" rather than "wine consumption" may be the basis of heart disease risk. In addition, traditional research is expensive and time-consuming, and many risk factors often cannot be randomly assigned. Therefore, we need more powerful methods to use observational data to evaluate causal relationships, and MR is such a method.
With the development of the human genome, people's understanding of the genetic factors of diseases has expanded from single-genes to multi-genes and multiple factors. It has been confirmed that many diseases are not determined by a single gene, but by the interaction of multiple genes, lifestyles, and environmental factors. For example, malignant tumors, coronary heart disease, diabetes, etc., may be affected by some changeable risk factors, such as diet, sleep, blood pressure, etc., in addition to genetic factors. The genome-wide association study (GWAS) has discovered a large number of genetic variations related to human diseases by testing the association between millions of genetic variations and disease outcomes, providing a basis for understanding disease mechanisms and predicting individual disease risk abilities. MR is a method that uses genetic data to evaluate the causal relationships between various risk factors, and these genetic variations also provide opportunities for MR.

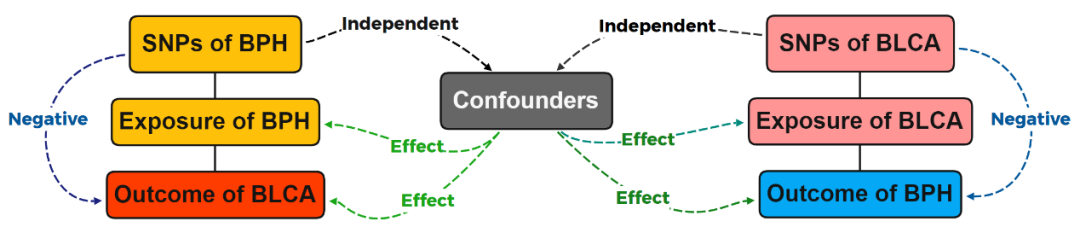
MR is an effective method for causal inference, which uses genetic variation as an instrumental variable to derive causal relationships between outcomes and exposures, effectively avoiding confounding bias in traditional epidemiological research. In addition, genetic information always spreads unidirectionally from parents to offspring, which limits the impact of reverse causality in MR.
This study explored the causal relationship between BPH and BLCA through the analysis of two samples of MR, enriched the pathogenesis of BLCA, and provided a new idea for the prognosis judgment and early clinical intervention of BLCA. The first author of this paper is Du Wenzhi, a doctoral student in the Department of Urology at Zhongnan Hospital of Wuhan University, supported by funding from the Hubei Provincial Department of Science and Technology.
References:
[1] Du W, Wang T, Zhang W, Xiao Y and Wang X (2022), Genetically supported causality between benign prostate hyperplasia and urinary bladder neoplasms: A mendelian randomization study. Front. Genet. 13:1016696. doi: 10.3389/fgene.2022.1016696.
Links:
https://www.frontiersin.org/articles/10.3389/fgene.2022.1016696/full